Tools & Resources
The Integration of Continuous Glucose Monitoring Data into the Electronic Health Record (iCoDE-1)
The role of the iCoDE consortium is to facilitate the efficient upload and integration of continuous glucose monitor (CGM) data into the Electronic Health Record (EHR). iCoDE aims to develop technical specifications to integrate CGM data into the EHR and develop workflows and guidelines to facilitate data integration efforts. Over 130 individuals from more than 60 organizations representing industry, academia, government agencies, clinicians, and patients participated in the year-long project. The result of this work is the 2022 iCoDE Report: CGM-EHR Integration Standards and Recommendations.
The Integration of Connected Diabetes Device Data into the Electronic Health Record (iCoDE-2)
The iCoDE-2 project is a consortium whose purpose is to facilitate efficient uploading and integration of insulin delivery data into the EHR. This builds on the 2022 iCoDE-1 foundation and extends the initiative to connected insulin delivery systems, including smart pens, pumps, and automated insulin delivery (AID) systems. iCoDE-2 sought to develop technical specifications to integrate insulin delivery data into the HER and develop workflows and guidelines to facilitate data integration efforts. The result of this work is the 2025 iCoDE Report: EHR integration Standards and Recommendations for Insulin Delivery Data.
Continuous Glucose Monitoring Logical Observation Identifiers Names and Codes (LOINC) Terms
Data standards and ontologies are critical for ensuring interoperability across information systems and can help drive adoption of novel data. Beyond standardization, these core elements should also be mapped to existing medical ontologies and specified by US Core Data for Interoperability (USCDI), such as Systematized Nomenclature of Medicine—Clinical Terms (SNOMED-CT), Logical Observation Identifiers Names and Codes (LOINC), Consolidated Clinical Document Architecture (C-CDA), and Observational Medical Outcomes Partnership (OMOP).
Given the critical role that standardized terminologies play in interoperability, data sharing, and research frameworks, we recommend that all elements of the iCoDE Core Dataset should have LOINC terms. We used the ATHENA – OHDSI Vocabularies Repository14 to search for existing terms that could be used. There are several limitations to the LOINC concepts, so we have developed new LOINC concepts that are being requested.
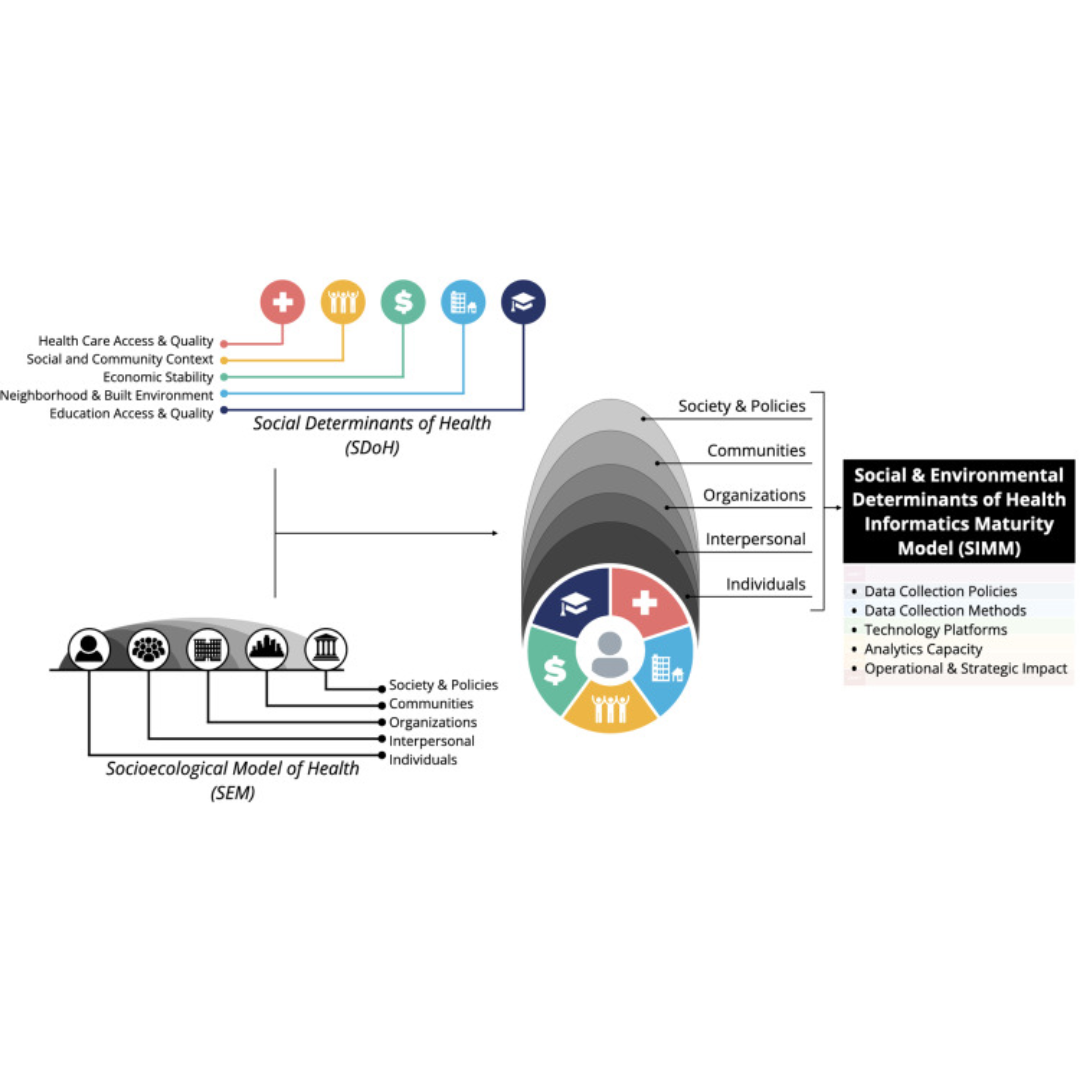
Social & Environmental Determinants of Health (SEDoH) Informatics Maturity Model
Integrating social and environmental determinants of health (SEDoH) into enterprise-wide clinical workflows and decision-making is one of the most important and challenging aspects of improving health equity. We engaged domain experts to develop a SEDoH informatics maturity model (SIMM) to help guide organizations to address technical, operational, and policy gaps.
We then developed a SIMM comprising seven maturity levels across five domains: data collection policies, data collection methods and technologies, technology platforms for analysis and visualization, analytics capacity, and operational and strategic impact. It’s designed to help organizations identify current gaps and next steps in improving SEDoH informatics. Improving the collection and use of SEDoH data is one important component of addressing health inequities.
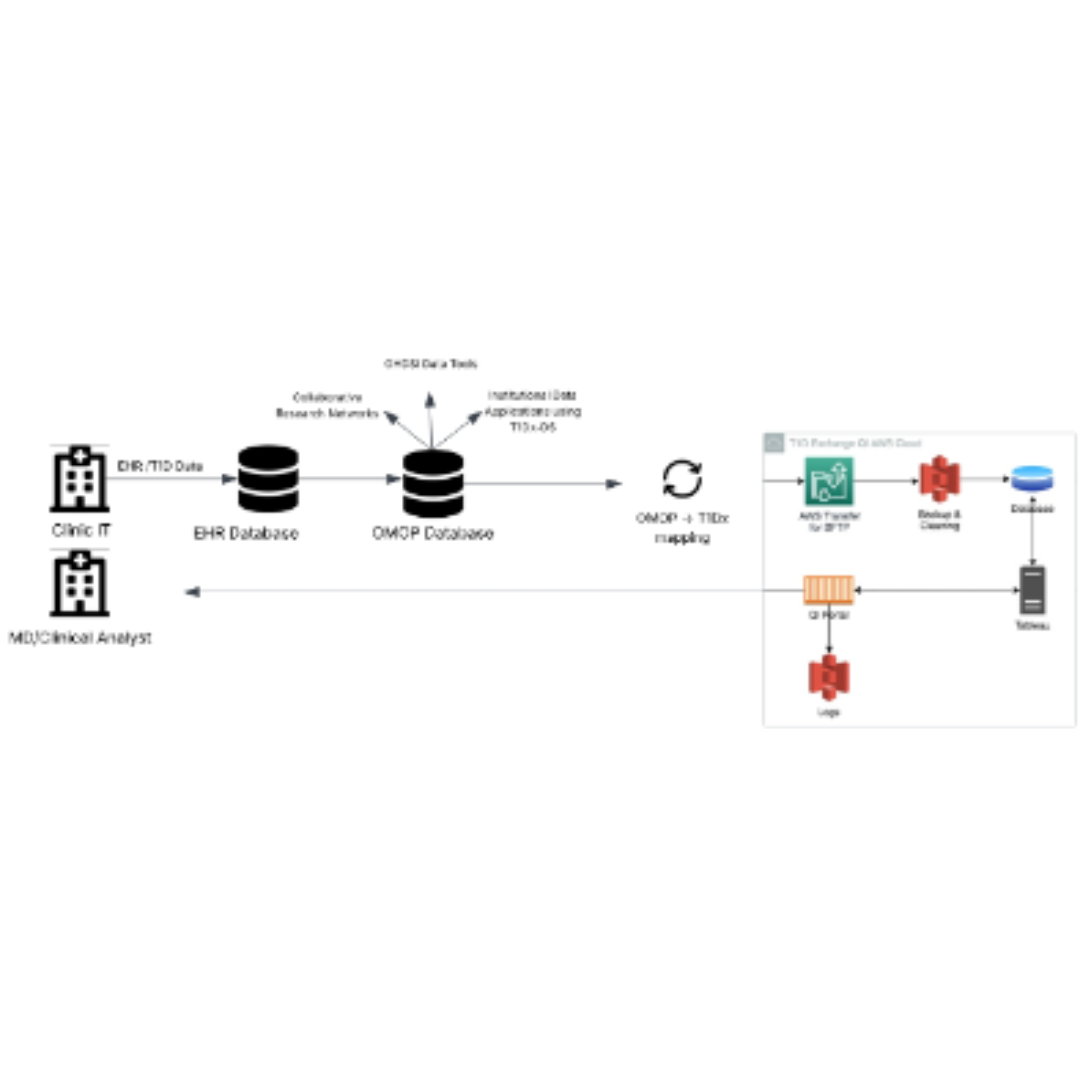
T1D Exchange (T1DX) OMOP Mapping
The T1Dx Data Specification (T1Dx-DS) has evolved over time and now includes more than 120 distinct variables, categorized into seven groups: Patients, Providers, Encounters, Observations, Conditions, Medications, and Diabetes.
Common Data Models (CDMs) are powerful tools that enable multi-organization collaborative research. The Observational Medical Outcomes Partnership (OMOP) CDM has become one of the most widely adopted CDMs in healthcare research. A number of robust, open-source tools already exist for OMOP, and many organizations already host their data in OMOP. Mapping the T1Dx-DS to OMOP would enable the T1Dx-QI to leverage these tools and infrastructure to accelerate research.
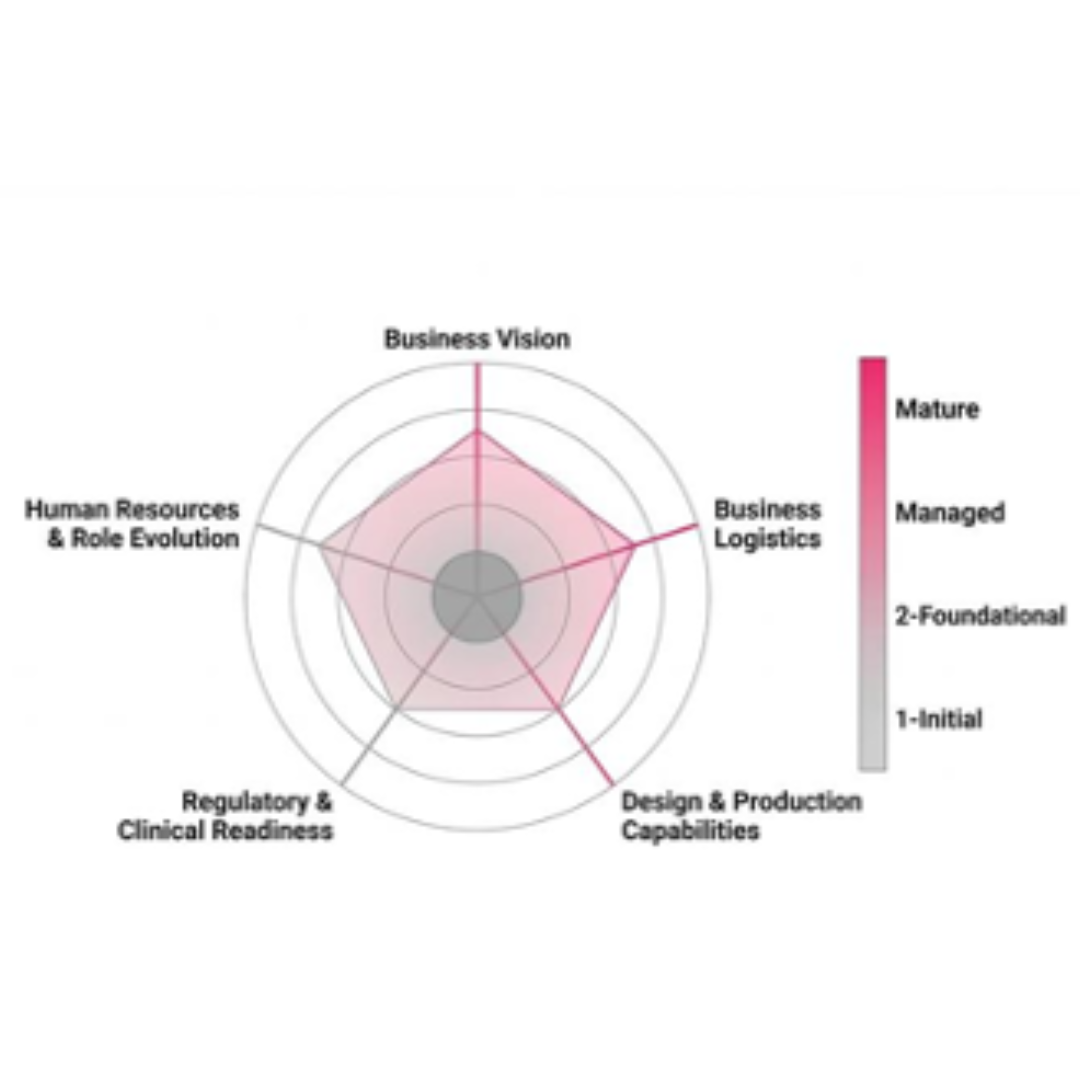
MedTech Startup Maturity Index (SMI)
Startup companies in the healthcare sector often fail because they lack sufficient entrepreneurial, regulatory, and business development expertise. Maturity models provide useful frameworks to assess the state of business elements more systematically than heuristic assessments. However, previous models were developed primarily to characterize the business state of larger, non-medical companies.
A novel MedTech Startup Maturity Index (SMI) was developed by a collaborative team of academic and industry experts and refined through feedback from external stakeholders. Pediatric medical device startups associated with the Consortium for Technology & Innovation in Pediatrics (CTIP) were scored and ranked according to the SMI following semi-structured interviews. This maturity index specifically for startup companies in the medical product sector is designed to help identify areas in which targeted interventions could assist business development.
Diabetes Technology Maturity Model (DTMM)
The Diabetes Technology Maturity Model (DTMM) is a structured framework that helps healthcare organizations evaluate their current capabilities, identify gaps, and build a clear roadmap for advancing diabetes technology integration. It supports better patient outcomes, streamlined workflows, and long-term sustainability. As tools like continuous glucose monitors (CGMs), insulin pumps, automated insulin delivery systems, and population health platforms become central to care, organizations often face inconsistent adoption, fragmented workflows, and underused data. DTMM offers a clear, evidence-informed roadmap to close these gaps, align teams, and maximize both clinical and financial impact.
Diabetes Research Model Edge Software
DRH Edge is a lightweight, downloadable desktop application that enables researchers to locally clean, process, standardize, and de-identify CGM datasets prior to repository submission. By allowing preprocessing to occur behind institutional firewalls, Edge supports HIPAA- and IRB-compliant workflows while ensuring data conform to DRH’s common data model and metadata standards before upload. This lowers technical and governance barriers to participation and accelerates data onboarding. Within the broader Diabetes Research Hub (DRH) ecosystem, Edge serves as the critical bridge between local research environments and the cloud repository, facilitating secure, FAIR-aligned data ingestion and enabling scalable, interoperable CGM data sharing across studies and institutions.
FoodSpace
FoodSpace is a community-driven tool and shared resource designed to accelerate food allergy research through collaboration, standardization, and secure data sharing. It serves as a central hub where investigators can discover studies, access reusable research artifacts, and soon will be able to explore curated datasets. Through its Study Directory, Resource Library, Data Catalog, and secure Trusted Research Environment, FoodSpace will support governed, reproducible analysis. By advancing a canonical, FAIR- and TRUST-aligned food allergy data model, FoodSpace will enable interoperability across studies and institutions. As both a platform and a community resource, it brings researchers together to share tools and expertise, accelerating discovery and translation to treatment.